Like many R packages, the Likert package in R is awesome, but it is particular about how you make you Likert plot data before you put it in the plot. I’ve gotten it down to nine steps.
Using Features of Likert Plot Data to Improve Plot Appearance
- The first six steps are a workaround to get it so that you can use the levels in factor variables in the Likert plot data as labels in the legend without getting any data errors.
- Steps seven and eight are to make it so the plot prints the full original Likert statements next to the bar by copying them the actual names of dataframe columns.
- The final step shows how I like to formulate my Likert plot code with options that uses the Likert plot data we created.
Download the data and code from my repository on Github.

Step 1: Create Minimal Dataset of Likert Plot Data
The minimal dataset of Likert plot data has only a study ID (numeric), and the Likert variables you need for your plot. The variable names should be short and easy to type, but unintuitive to the statement (e.g., Q1, Q2, Q3, etc.). All variables should be in numeric format.
survey1 <- read.csv("survey_data.csv", header = TRUE, sep = ",")
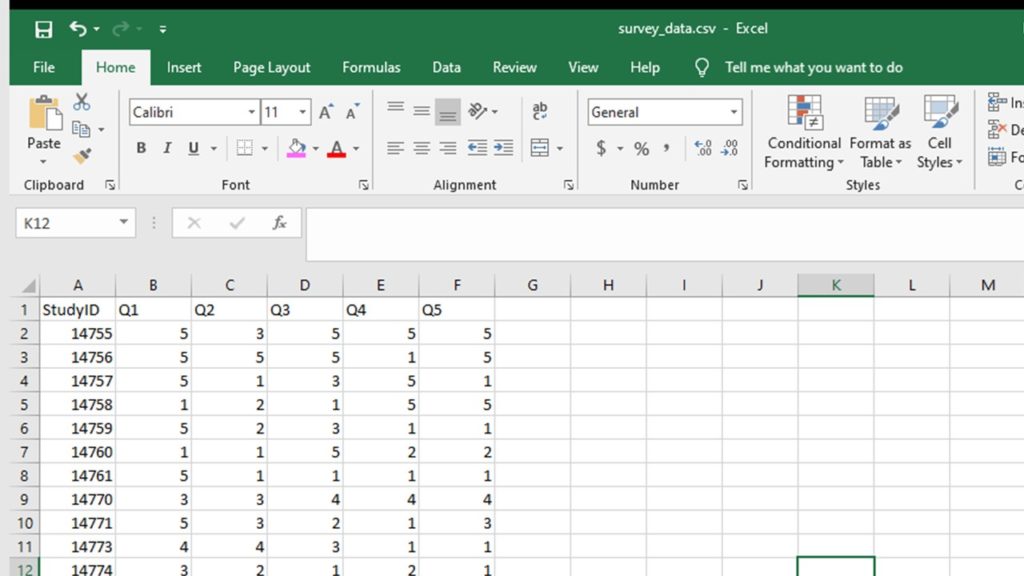

Step 2: Prepare to make a fake dataset, append it to the real dataframe, do some processing, then remove the fake records.
This is the hack to make it so the Likert plot works even if one or more of your statements has a level that wasn’t said by a respondent.
Imagine you have a Likert scale of
- 1 = Strongly Disagree
- 2 = Somewhat Disagree
- 3 = Neither Agree nor Disagree
- 4 = Somewhat Agree, and
- 5 = Strongly Agree.
Imagine you had a statement everyone agreed with – so in the dataset, there are only answers 4 and 5 to that statement. That means that levels 1, 2, and 3 are not represented, and this would create a complication when plotting. This is the hack to prevent this.

Step 3: Create fake dataset of Likert plot data in the same format as the minimal dataset with a row for each level of the Likert scale.
With our example scenario, it means we have five levels, so the fake dataset will have five rows. Each row should have a study ID within a known range (e.g. between 9991 and 9995) that is not in the range of the real study IDs, so you can remove these fake data rows later. Each row of the Likert plot data should be all coded with one of the levels. So, for us, we’d have row 9991 and make every answer be 1, then a row for 9992 and make every answer be 2, etc.
StudyID <- c("99991", "99992", "99993", "99994", "99995")
Q2 <- c(1, 2, 3, 4, 5)
Q3 <- c(1, 2, 3, 4, 5)
Q4 <- c(1, 2, 3, 4, 5)
Q5 <- c(1, 2, 3, 4, 5)
fake <- data.frame(StudyID, Q1, Q2, Q3, Q4, Q5)
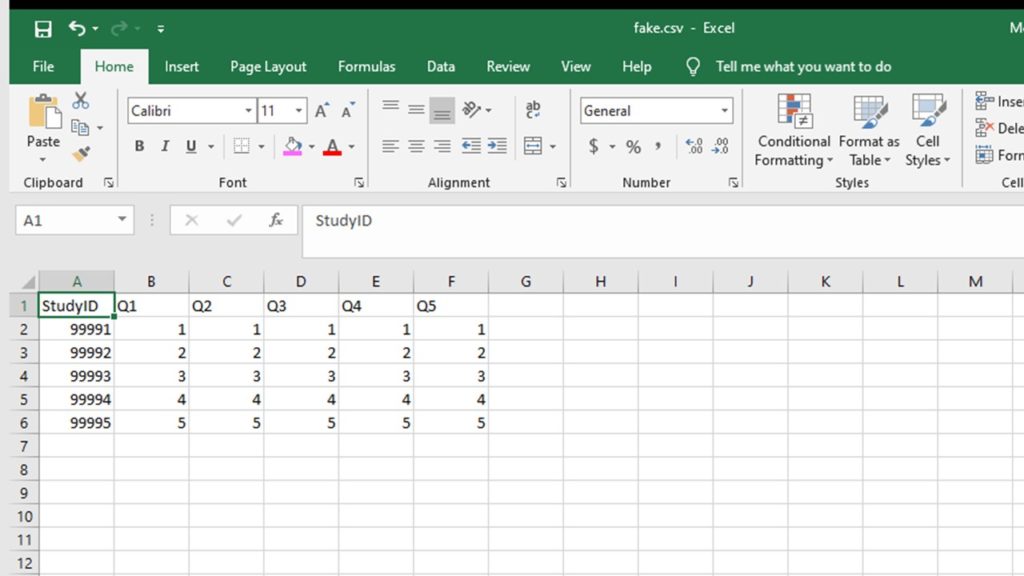

Step 4: Append the fake dataframe to the real dataframe, and convert all of the variables to factor format.
Factor data format is really specific to R. It’s meant for codes for ordinal or nominal variables (exactly what we are doing). That way, R sees the values not as numbers, but as codes, and can attach labels to each code; these labels are called “levels”. See this information page from UC Berkeley for more on the factors in R concept. For SAS users, the parallel is attaching formats to numerical variables using PROC FORMAT. Now that we have appended the fake data on, and every level is represented in the dataset for each Likert variable, we can convert them all to factor. I like to put them all in new variables with the _f suffix (e.g., Q1_f, Q2_f, Q3_f).
survey2 <- rbind(survey1, fake) survey2$Q1_f <- as.factor(survey2$Q1) survey2$Q2_f <- as.factor(survey2$Q2) survey2$Q3_f <- as.factor(survey2$Q3) survey2$Q4_f <- as.factor(survey2$Q4) survey2$Q5_f <- as.factor(survey2$Q5)

Step 5: Attach factor levels to each of the Likert statement variables you just converted to a factor.
As I said above, the factor levels are essentially labels for each level in the Likert plot data. Because we added the fake data, we can be sure that none of the levels are missing in any of the data, and the levels will get properly attached.
factor_levels <- c("Strongly Disagree","Somewhat Disagree","Neither Agree nor Disagree",
"Somewhat Agree","Strongly Agree")
levels(survey2$Q1_f) <- factor_levels
levels(survey2$Q2_f) <- factor_levels
levels(survey2$Q3_f) <- factor_levels
levels(survey2$Q4_f) <- factor_levels
levels(survey2$Q5_f) <- factor_levels

Step 6: Remove the fake data.
This is easily done by filtering a saving a new dataframe without the rows with the study IDs assigned to the fake data (in our case, rows 9991 to 9995).
nrow(survey2) survey3 <- subset(survey2, StudyID < 99991) nrow(survey3)

Step 7: Remove the study ID column, and all the original numeric columns without the _f suffix.
In other words, make a plotting dataframe of just the variables with the _f suffix. This is because we need the entire dataset to have only the factor format Likert variables with the levels attached, or the plot doesn’t work.
colnames(survey3) survey4 <- survey3[,7:11] colnames(survey4)

Step 8: Replace the Likert plot data column headings with the actual Likert statement
This is easiest done by making a vector, and then replacing the dataframe headings with the vector. Now you are done with your Likert plot data!
VarHeadings <- c("I want to live in a world with unicorns.",
"Whenever given a choice, I choose chocolate.","My hair is too long.",
"There really aren't any reasons to do cross-stitch.","Rats are misunderstood.")
names(survey4) <- VarHeadings
colnames(survey4)

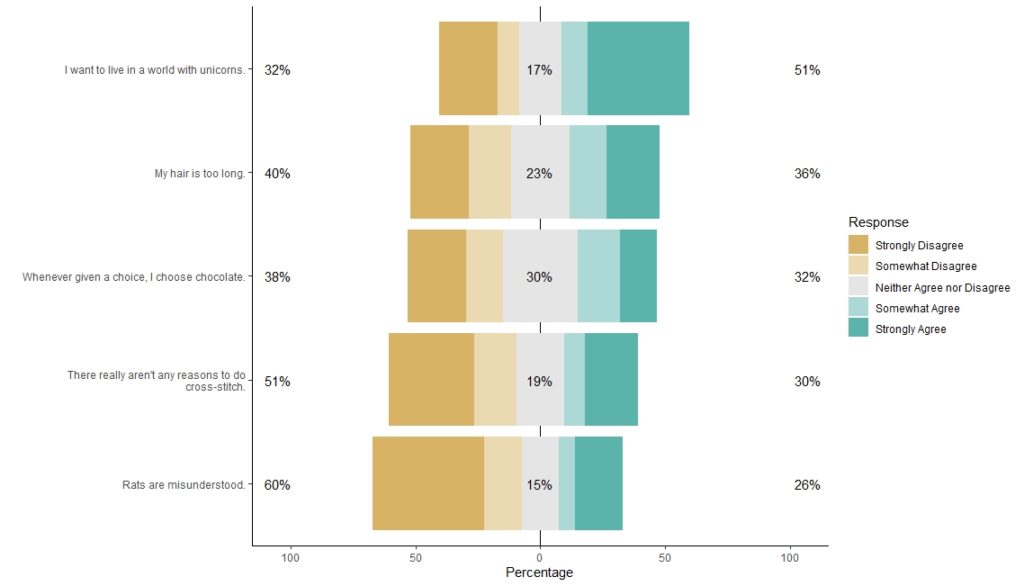
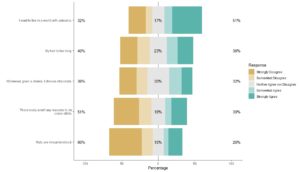
Step 9: Plot the dataset with a legend.
Through our machinations, the plot is labeled clearly with the Likert statements, and the legend is labeled clearly with the levels.
library(likert) p <- likert(survey4) a <- likert.bar.plot(p, legend.position = "right", text.size = 4) + theme(text = element_text(size = rel(4)),axis.text.y = element_text(size = rel(2))) + theme_update(legend.text = element_text(size = rel(0.7))) + theme_classic() plot(a)
Updated October 10, 2020. Added FTC disclaimer and other edits on December 5, 2020. Formatting edits September 12, 2021 and October 11, 2022. Added banners March 6, 2023. Revised banners June 19, 2023. Revised banners June 24, 2024.
Read all of our data science blog posts!
I love the Likert package in R, and use it often to visualize data. The problem is that sometimes, I have sparse data, and this can cause problems with the package. This blog post shows you a workaround, and also, a way to format the final plot that I think looks really great!





















































This was excellent! Thank you for being my collaborator. I really wanted to use this package and was getting so frustrated with how to make it work; adding in the fake data did the trick 🙂
Awesome! If you go to the video, you will see one of the commenters wrote some more complex code to solve the problem. I’m not smart enough for that – so adding fake data does the trick!